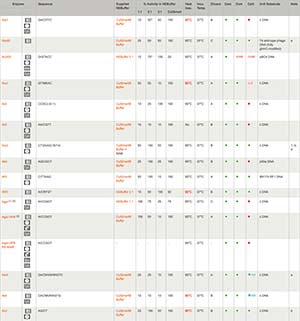 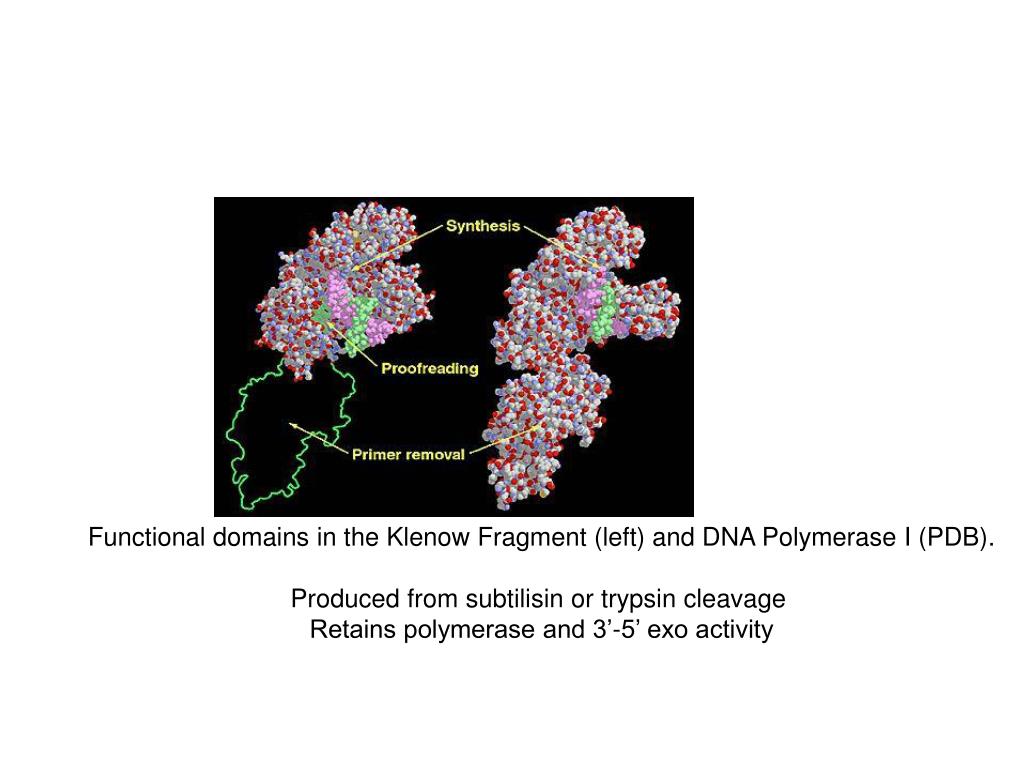
Isolation and extraction of blunted vector fragment- I gel purified my fragment by freeze-thaw, ethanol precipitated the DNA, and resuspended in 20 uL CIAP buffer (NEB 3). I then heat inactivated Klenow by heating at 75 C for 20 minutes. I just wanted to be on the super-safe side.įill-in Reaction- I added 2 uL of Klenow (NEB: 3'-5' exo-) and 1 uL of 2.5 mM dNTPs and incubated at 37 C for 30 minutes. In fact, it really wouldn't have mattered if I didn't heat inactivate them because the blunting reaction wouldn't have recreated their recognition sites. I was lucky that both NdeI and BsrGI can be heat inactivated. I inactivated the restriction enzymes by first upping the salt to 100mM and then heating at 80 C for 20 minutes. I incubated the reaction for 2 hours at 37 C and ran 1 uL to check for completion of the digest. Preparation of vector- I cut my vector DNA with NdeI and BsrGI in a 50 uL reaction containing NEB 4. Also, I didn't care which way my fragment my fragment went in. I would avoid doing a blunt end ligation if you can find compatible enzymes or can add handy cleavage sites by PCR. *Considerations* There simply wasn't another way to achieve the right construct without doing it this way. I read everything I could about the topic on this forum and came up with the following protocol, which, worked really well. I have recently performed a successful ligation using Klenow to make blunt ends. Ones 1 in 72 colonies and many times I just have nothing !! I wish someone could give me some good advices or show me how to do it with better results! Ones, I had 5 positive colonies in 48 I picked. Ones of them, incubating 15 min, the other one, 4 or 6 h at 25Â☌ both (I inactive the first one at 70Â☌ 20 min to wait for the second one). Two tubes kenow (NEB) treatment using twice that NEB recommends but no more in any NEB restriction buffer and dNTPs at 50 uM. Gel-purification (if I don’t do that, I have some or many colonies in my c(-). Digest my DNA (a good amount), phenol-chlorof extraction (I think it wouldn’t be necessary, but I do as my boss recommends). What I have done with the best results is: I have try many different conditions!!!! Temperature, klenow concentration, incubation time. She says it’s very important a Phenol-Chloroform extraction before and after klenow treatment and she says she incubated at 37Â☌ over night. I have try it many times with very few successful cases.īy boss says she did it many times (I don’ t know). I’m suffering a lot because I have to fill in many different thing (The reason? A long story ). I’m going to talk about my experience…. Libraries were quantified with the Bioanalyzer 2100 (Agilent) and sequenced on the Illumina HiSeq 2000 Sequencer (100 nucleotide read length).I don’t think I could help you very much. PCR products were size selected using 3% NuSieve agarose gels (Lonza) followed by gel extraction using QIAEX II reagents (QIAGEN). 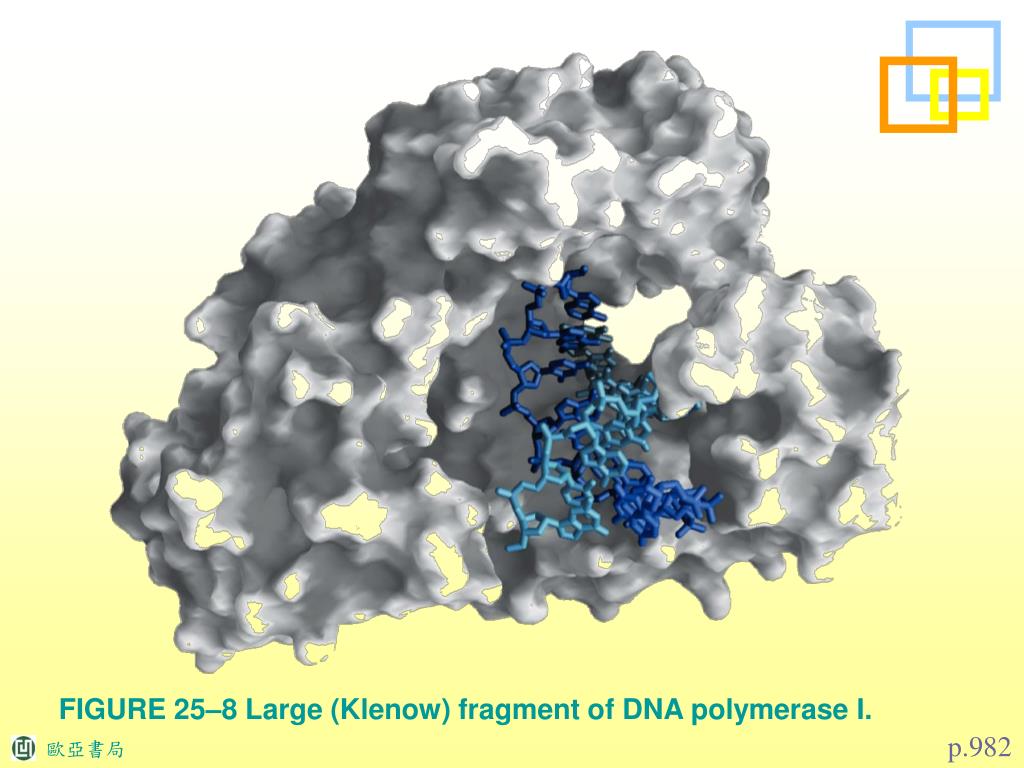
The adaptor-modified DNA fragments were enriched by PCR using the Illumina Barcode primers and Phusion DNA polymerase (NEB). A single A-base was added to fragment ends by Klenow fragment (3' to 5' exo minus NEB) followed by ligation of Illumina adaptors (Quick ligase, NEB). ChIP-enriched DNA samples (1-10 ng) were converted to blunt-ended fragments using T4 DNA polymerase, E.coli DNA polymerase I large fragment (Klenow polymerase) and T4 polynuleotide kinase (New England BioLabs, NEB). The ChIP-seq sample preparation for sequencing was performed according to the manufacturer's instructions (Illumina). These samples were then immunoblotted as described above with the exception of using protein A-HRP secondary (GE HealthCare, Cat#: NA9120-1ML) antibody for detection. Beads were washed 3 times with IP buffer (150nM NaCl Thermo Fisher Scientific) and directly boiled in 1X NuPage LDS/reducing agent buffer (ThermoFisher Scientific Cat#: NP0007 and NP0009) to elute and denature the precipitated proteins. Next day, 50ul of Dynabeads Protein-G beads were added to the lysate-antibody mixture and incubated for 2h at 4C. A fraction of the cleared lysate was saved as input and the remainder was incubated overnight (12-16 hours) with 10ug of target protein antibody at 4C with gentle mixing. Nuclear lysates were incubated for 2 hours at 4C with 30ul of magnetic Protein-G Dynabeads (Thermo Fisher Scientific Cat#: 10004D) for pre-clearing. Library_strategy ChIP-Seq library_source GENOMIC library_selection ChIP library_construction_protocol 8-10 million cells ectopically overexpressing different V5-tagged FOXA1 variants and WT AR (or TLE3) were fractionated to isolated intact nuclei using the NE-PER kit reagents (Thermo Fisher Scientific Cat#: 78835) and lysed in the complete IP lysis buffer (Thermo Fisher Scientific Cat#: 87788).
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
